 |
DNA Microarrays: A Case Study
The great challenge in DNA microarrays problems consists in finding the genes
that are relevant to a particular disease or cell state in the midst of thousands
of other genes. And thinking of genes as variables, for tackling these problems
successfully, in an ideal world, one would have required about an order of magnitude
more of samples. However, in a typical DNA microarray study, the number of samples
available (a few dozens or a few hundreds at best), when compared to the thousands
of genes under study, is ridiculously small, which obviously poses some challenges.
First, it is common to try and narrow down the search space with sophisticated
discretization algorithms to filter out the noise or irrelevant genes. And although
good results have been reported (data size reductions of 50%-98%), the problem still
remains: hundreds or thousands of variables to be mined using just a few dozen samples.
The great advantage of using GeneXproTools in DNA microarray studies is that you can
use the raw data (obviously you can also use the filtered data and even use GeneXproTools
to filter out the noise) and still obtain excellent results. Obviously this means that not
all the models with a good accuracy on the training data will have good predictive accuracy,
but by making several runs one can select the top 10-20 models and then cross-reference the genes
(attributes) used in all of them. You can then select and copy these most important genes from the
Data Panel and create a much smaller dataset for creating the final model.
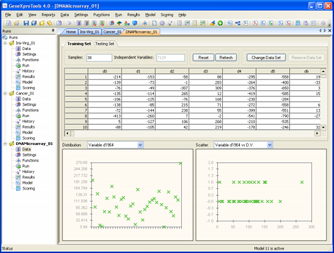
Let's illustrate this with real-world DNA microarray data, using the
well-studied ALL-AML Leukemia datasets (these
same datasets are used in the DNA Microarray sample run of
GeneXproTools 4.0, and we recommend you play with it as you'll be
able to see everything including the generated code). In this
problem, the training dataset consists of 38 bone marrow samples (27 ALL and 11 AML), over 7129 probes from 6817 human genes.
And the testing dataset consists of 34 samples, with 20 ALL and 14 AML.
For this analysis, "0" was used to represent "ALL" and "1" to
represent "AML" and the 7129 genes were numbered d0-d7128.
For instance, in one study, for the
ALL-AML Leukemia problem, a total of 308 promising genes were
identified in 25 good runs (that is, in this case runs with 100% training
accuracy and testing accuracies between 91.18%-97.06%). Of these 308
promising genes, only 11 (genes 759, 1881, 2287, 2407, 4362, 4846, 5485, 6040, 6587, 6638,
and 6854) appeared in two or more models; and of these, only five (genes
759, 1881, 2287, 4846, and 6854) appeared in more than three models,
with the most prevalent being genes 1881, 2287, and 4846, with
eight, six, and four appearances, respectively. So, it is a good
guess that the genes mostly to be involved in ALL-AML leukemia are
genes 1881, 2287, and 4846, which is an exceptionally good starting
point for tackling leukemia.
References
Golub, T. R., D. K. Slonim, P. Tamayo, C.
Huard, M. Gaasenbeek, J. P. Mesirov, H. Coller, M. L. Loh, J. R.
Downing, M. A. Caligiuri, C. D. BloomÞeld, and E. S. Lander, 1999.
Molecular Classification of Cancer: Class Discovery and Class Prediction by Gene Expression
Monitoring. Science, 286:531-537.
Last modified: September 30, 2006
Cite this as:
Ferreira, C. "DNA Microarrays: A Case Study." From GeneXproTools
Tutorials – A Gepsoft Web Resource.
https://www.gepsoft.com/tutorial001.htm
|

